Illumina – Shining a Light on Your DNA

How technological advances in gene sequencing is making personalized healthcare increasingly affordable.
In 2003, an international scientific initiative called the Human Genome Project mapped the first complete human genome. The cost was incredible – $3 billion and 15 years of research – but it was worth it. Identifying the complete sequence of the human genome was akin to discovering the operating manual of the human body. In essence, the project proved a major milestone to a nascent field, genomics, which is dedicated to studying how our genetic make-up affects our health needs.
Since each human body has its own unique manual, individualized analysis is necessary for genomics-based research to truly prosper and impact preventative, personalized medicine. But the cost in 2003 ($300 million per run) made that dream infeasible.
In the past 15 years, Illumina has broken Moore’s Law in the cost of gene sequencing through a laser focus on digital innovation in a new method called “next-generation sequencing” (NGS). Using big data analysis and new techniques that performs millions of sequencing reactions concurrently, Illumina’s operating model has created a paradigm shift in personalized DNA services using the machines it develops.

To get a sense of how powerful digital transformation is on genomics, in 2005, a single sequencing run could only produce one gigabase (one billion base pairs) of data and cost $14 million. By late 2014, that rate had climbed to 1.8 terabases of data (1,800x increase) while the cost has come down to $4,000 (3,500x decrease). And just this year, Illumina’s HiSeq X machine broke $1,000.
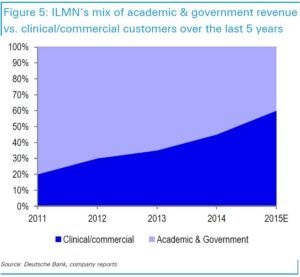
To date, only under 500,000 people have had their genomes sequenced – the fixed costs (over $5 million for a high throughput machine) are still too prohibitive even for the $1,000 run. And despite Illumina owning >90% of the NGS market, academia and government research still represent about half the revenues. But as costs continue to decrease, volume should grow along with the total addressable market, which has now expanded to prenatal care and oncological diagnostics. Illumina’s business model is poised for this virtuous cycle due to digital transformation: the cheaper the technology becomes, the more likely it will be reimbursed by insurance, the better the data set becomes, leading to new peripheral fields if costs could further decline.
Digital transformation is at the heart of Illumina’s operating model, but its ability to attract human and investment capital comes from its business model. Despite being a leader in a still nascent technology, Illumina is cash flow positive – with $2.4 billion in 2016E revenues and $500 million in free cash flow. Operating margins are 35%, despite having the industry’s highest R&D budget at 20% of sales. Illumina is a rare breed of startup – a profitable, tangible business that has high recurring revenue and a propensity to internally reinvest. Its business model of two-thirds consumables and one-third equipment sales allow for this dynamic. Each new Illumina customer who buys its machines will typically spend 30-40% of the machine’s value in consumables each year. These consumables are proprietary to Illumina, so once the lock (equipment) is sold, only an Illumina manufactured key (consumable) can be used.
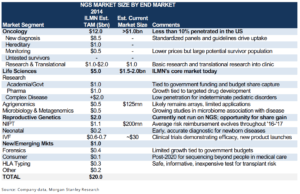
Perhaps what is even more exciting about this business model is an ever-expanding total addressable market. In 2014, Illumina quantified its TAM at $20 billion, but one of the greatest impact of big data gene sequencing is an emerging field called population genomics, or essentially big data analysis on NGS. Population genomics is the large-scale comparison of DNA across populations to study the genome-wide effects. These genetic studies yield important clues on locating genes underlying complex diseases, and is opening new applied fields from forensics to archaeology to transplant genomics (which helps determine the compatibility of a biological donor and recipient).
Going forward, I believe that Illumina should fully embrace vertical integration downstream into the currently fragmented market of consumer genomics (e.g., 23andMe, Ancestry.com, etc.). The next digital transformation in this industry should be the creation of vast genomic libraries. As more individuals sequence their genes, Illumina should move away from only equipment manufacturing and become the Google of genomics. So far, management has only dipped a toe with an equity investment in Helix. But if Illumina were to wholeheartedly embrace consumer genomics, it can sequence and store consumers’ DNA, then sell through an app platform offered by third-party players. Helix has already signed on partners such as LabCorp and Mayo Clinic, but a full-fledged push downstream with a continued emphasis on big data analytics could create the world’s largest sequencing laboratory.
Next generation sequencing is opening our lives to a very powerful form of preventative medicine. Imagine an ecosystem of physicians, nurses, and genetic counselors who personalize your healthcare experience. And through new analytic like population genomics, we can understand at a molecular level how illnesses come about, and discover customized drugs that minimize side effects in a way unavailable today.
798 WORDS

Research resources:
- Illumina Company Presentations, IR website, Annual Reports, and other SEC Filings
- National Institute of Health
- Analyst reports from Deutsche Bank, Morgan Stanley, Cowen & Company, and Goldman Sachs
- http://users.ugent.be/~avierstr/nextgen/Next_generation_sequencing_web.pdf
- http://www.ebi.ac.uk/training/online/course/ebi-next-generation-sequencing-practical-course/what-you-will-learn/what-next-generation-dna-
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3841808/
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4254436/
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3080771/
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3186121/
- https://www.technologyreview.com/s/540711/inside-illuminas-plans-to-lure-consumers-with-an-app-store-for-genomes/



Thanks for the article Lawrence.
I actually used Illumina to do gene analysis on mice cells in my thesis on hair follicle regeneration, so I know how useful next gen sequencing is for academic research. As NGS cost continues to fall, I think we’ll see NGS more widely applied for personal genomics. If the 23&Me’s of the world can convince consumers to be interested in their genome, Illumina will be able to access more data, as you said. But, for consumers who want NGS to help their doctors develop drugs for their disease, we cannot stop at NGS.
The big bottleneck is not in genomic analysis these days, but in drug development. Even if we know what genes are mutated in various diseases, it is still a very time consuming and costly process to generate chemical compounds that can turn into effective drugs. While efforts are being made to speed up this process (particularly at the Broad Institute of Harvard/MIT), we may still be a decade away from seeing the true benefits of NGS on actual patients.
Interesting article and perspective. I think the knowledge that could be gained from massive sequenced genome libraries would be massively beneficial to our understanding up modern diseases and genetic disorders. I personally see the largest challenge in encouraging the general population to get their genome sequenced and the data aggregated. In the modern day where privacy is hard to come by, I see many people not wanting their personal information, health history, and genetic propensity towards disease aggregated by any company or government organization. I believe that to aggregate this data, companies or organizations will have to strongly encourage the masses to submit to this testing and sequencing and until that happens, I believe these libraries will be grossly underpopulated and, to some extent, ineffective at truly mapping out genetic connections between diseases and disorders.
Extremely interesting article! I agree with you that genome sequencing and personalized medicine are the future of healthcare, and Illumina holds a very valuable position as the market leader in NGS technology. I just want to point out one risk in shifting focus away from academic/clinical customers in favor of commercial customers as you suggest. Our ability to generate genomic data is moving at a much faster pace than our ability to interpret it (when is a genetic mutation actually significant clinically?) as well as to do something about it, as Paul points out above. This discrepancy has made many stakeholders in the healthcare field, including regulators, hesitant about placing a large amount of difficult to interpret data in the hands of consumers. For example, 23andme recently struggled with FDA approval, was forced to shut down for a period, and ultimately was allowed to return to market but with a product that was much more limited in scope.
I agree with all the post above. This is an awesome write-up. I do have to agree with MattGerow. While I think the technological innovation here is second to none there is a key barrier to entry here. Will people be willing to part with information that they believe to be so personal? It seems unlikely to me now but the potential of the company’s technology is too significant to ignore.
Thanks for a really interesting post. The decrease in cost from 2003 at $300M per run to $1000 in 2016 is stunning and speaks to the potential for future applications of this technology. The potential applications sound very appealing in their ability to identify and diagnose health problems and even tailor treatment to individual patients. However, I do wonder what the potential negative consequences would be if this type of genomic sequencing were to become more common. What would be the consequences of the availability of data describing my specific DNA? How could I be judged, perhaps in contexts outside of medicine, based on my genetic characteristics? The benefits associated with the individualized care and population genomics may outweigh the negative consequences, but those possible negative consequences of making that DNA information more readily accessible is what gives me pause. There needs to be strict data protection guidelines and ethical thought that proceeds in parallel with these advancements.